Copyright notice:
The material is presented to ensure timely dissemination
of scholarly and technical work. Copyright and all rights
therein are retained by authors or by other copyright holders.
All persons copying this information are expected to
adhere to the terms and constraints invoked by each
copyright. In most cases, these works may not be
reposted without the explicit permission of the
copyright holder.
If you want a paper that is not available here,
or if you have any problems accessing these documents,
e-mail me.
MatureBayes
MicroRNAs (miRNAs) are small, single stranded RNAs with a key
role in post-transcriptional regulation of thousands of genes
across numerous species. While several computational methods
are currently available for identifying miRNA genes, accurate
prediction of the mature miRNA remains a challenge. Existing
approaches fall short in predicting the location of mature
miRNAs but also in finding the functional strand(s) of miRNA
precursors. MatureBayes is a computational tool that incorporates
a Naive Bayes classifier to identify mature miRNA candidates
based on sequence and secondary structure information of
their miRNA precursors. We take into account both positive
(true mature miRNAs) and negative (same-size non-mature
miRNA sequences) examples to optimize sensitivity as well as
specificity. Our method can accurately predict the start
position of experimentally verified mature miRNAs for both
human and mouse, achieving a significantly larger (often double)
performance accuracy compared with two existing methods. Moreover,
the method exhibits a very high generalization performance
on miRNAs from two other organisms. More importantly, our
method provides direct evidence about the features of miRNA
precursors which may determine the location of the mature miRNA.
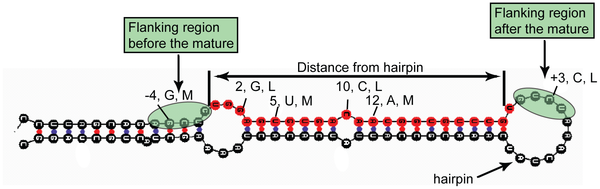
We find that the triplet of positions 7, 8 and 9 from the
mature miRNA end towards the closest hairpin have the largest
discriminatory power, are relatively conserved in terms of
sequence composition (mostly contain a Uracil) and are located
within or in very close proximity to the hairpin loop,
suggesting the existence of a possible recognition site for
Dicer and associated proteins
Publications
-
MatureBayes: a probabilistic algorithm for
identifying the mature miRNA within novel
precursors.
Gkirtzou Katerina, Tsamardinos Ioannis, Tsakalides Panagiotis and Poirazi Panagiota.
PLoS One, 2010
[pdf] [bibtex] -
Mature miRNA identification via the use of a
Naive Bayes Classifier
Gkirtzou Katerina
Master of Science, University of Crete, 2009
[pdf] [bibtex] [presentation] -
Mature miRNA Identification via the Use of a
Naive Bayes Classifier.
Gkirtzou Katerina, Tsakalides Panagiotis and Poirazi Panagiota.
8th IEEE International Conference on BioInformatics and BioEngineering (BIBE), 2008
[pdf] [bibtex]
Code/Tool
The code is provided under the GNU GPL license. It does not come with any warranty of any kind.Web interface is available here.
Code is available here.